Checking your network
Selecting which rates to include and which to exclude from your network is a bit of an art. pynucastro has a few tools to help check what you might be missing.
[1]:
import pynucastro as pyna
[2]:
reaclib_library = pyna.ReacLibLibrary()
RateCollection validate method
Let’s start by trying to create a network for carbon burning.
To start, let’s pick the nuclei \(\alpha\), \({}^{12}\mathrm{C}\) and \({}^{20}\mathrm{Ne}\)
[3]:
nuclei = ["he4", "c12", "ne20"]
cburn_library = reaclib_library.linking_nuclei(nuclei)
Now we can make a rate collection
[4]:
rc = pyna.RateCollection(libraries=[cburn_library])
rc
[4]:
C12 ⟶ He4 + He4 + He4
C12 + C12 ⟶ He4 + Ne20
Ne20 + He4 ⟶ C12 + C12
He4 + He4 + He4 ⟶ C12 + 𝛾
Now, since we are primarily interested in \({}^{12}\mathrm{C} + {}^{12}\mathrm{C}\), let’s make sure we are not missing any other reactions that have the same reactants. The validate() method will do this, by comparing the rates we have selected to another library.
[5]:
rc.validate(reaclib_library)
validation: Ne20 produced in C12 + C12 ⟶ He4 + Ne20 never consumed.
validation: missing He4 + He4 + He4 ⟶ p + B11 as alternative to He4 + He4 + He4 ⟶ C12 + 𝛾 (Q = -8.682 MeV).
validation: missing He4 + He4 + He4 ⟶ n + C11 as alternative to He4 + He4 + He4 ⟶ C12 + 𝛾 (Q = -11.4466 MeV).
validation: missing C12 + C12 ⟶ p + Na23 as alternative to C12 + C12 ⟶ He4 + Ne20 (Q = 2.242 MeV).
validation: missing C12 + C12 ⟶ n + Mg23 as alternative to C12 + C12 ⟶ He4 + Ne20 (Q = -2.598 MeV).
[5]:
False
This tells us that we are missing 2 branches of the \({}^{12}\mathrm{C} + {}^{12}\mathrm{C}\) reaction. ReacLib already scales the rates based on the branching of the products, so we should try to include these other branches.
Note: by default, validate() only checks forward rates.
To address these issues, we need to include the additional nuclei. In particular, the branch that makes \({}^{23}\mathrm{Na}\) is likely important (the rate making \({}^{23}\mathrm{Mg}\) is endothermic, so less likely).
[6]:
nuclei += ["p", "na23"]
cburn_library = reaclib_library.linking_nuclei(nuclei)
rc = pyna.RateCollection(libraries=[cburn_library])
rc
[6]:
C12 ⟶ He4 + He4 + He4
C12 + C12 ⟶ p + Na23
C12 + C12 ⟶ He4 + Ne20
Ne20 + He4 ⟶ p + Na23
Ne20 + He4 ⟶ C12 + C12
Na23 + p ⟶ He4 + Ne20
Na23 + p ⟶ C12 + C12
He4 + He4 + He4 ⟶ C12 + 𝛾
[7]:
rc.validate(reaclib_library)
validation: Ne20 produced in C12 + C12 ⟶ He4 + Ne20 never consumed.
validation: Ne20 produced in Na23 + p ⟶ He4 + Ne20 never consumed.
validation: missing He4 + He4 + He4 ⟶ p + B11 as alternative to He4 + He4 + He4 ⟶ C12 + 𝛾 (Q = -8.682 MeV).
validation: missing He4 + He4 + He4 ⟶ n + C11 as alternative to He4 + He4 + He4 ⟶ C12 + 𝛾 (Q = -11.4466 MeV).
validation: missing C12 + C12 ⟶ n + Mg23 as alternative to C12 + C12 ⟶ He4 + Ne20 (Q = -2.598 MeV).
validation: missing C12 + C12 ⟶ n + Mg23 as alternative to C12 + C12 ⟶ p + Na23 (Q = -2.598 MeV).
validation: missing Na23 + p ⟶ n + Mg23 as alternative to Na23 + p ⟶ He4 + Ne20 (Q = -4.839 MeV).
validation: missing Na23 + p ⟶ Mg24 + 𝛾 as alternative to Na23 + p ⟶ He4 + Ne20 (Q = 11.6927 MeV).
[7]:
False
Now, looking at what is missing, we probably want to include \({}^{24}\mathrm{Mg}\) as an endpoint for carbon burning.
[8]:
nuclei += ["mg24"]
cburn_library = reaclib_library.linking_nuclei(nuclei)
rc = pyna.RateCollection(libraries=[cburn_library])
rc
[8]:
Mg24 ⟶ p + Na23
Mg24 ⟶ He4 + Ne20
C12 ⟶ He4 + He4 + He4
Ne20 + He4 ⟶ Mg24 + 𝛾
Na23 + p ⟶ Mg24 + 𝛾
C12 + C12 ⟶ p + Na23
C12 + C12 ⟶ He4 + Ne20
Ne20 + He4 ⟶ p + Na23
Ne20 + He4 ⟶ C12 + C12
Na23 + p ⟶ He4 + Ne20
Na23 + p ⟶ C12 + C12
He4 + He4 + He4 ⟶ C12 + 𝛾
[9]:
rc.validate(reaclib_library)
validation: Mg24 produced in Ne20 + He4 ⟶ Mg24 + 𝛾 never consumed.
validation: Mg24 produced in Na23 + p ⟶ Mg24 + 𝛾 never consumed.
validation: missing He4 + He4 + He4 ⟶ p + B11 as alternative to He4 + He4 + He4 ⟶ C12 + 𝛾 (Q = -8.682 MeV).
validation: missing He4 + He4 + He4 ⟶ n + C11 as alternative to He4 + He4 + He4 ⟶ C12 + 𝛾 (Q = -11.4466 MeV).
validation: missing C12 + C12 ⟶ n + Mg23 as alternative to C12 + C12 ⟶ He4 + Ne20 (Q = -2.598 MeV).
validation: missing C12 + C12 ⟶ n + Mg23 as alternative to C12 + C12 ⟶ p + Na23 (Q = -2.598 MeV).
validation: missing Ne20 + He4 ⟶ n + Mg23 as alternative to Ne20 + He4 ⟶ Mg24 + 𝛾 (Q = -7.21457 MeV).
validation: missing Na23 + p ⟶ n + Mg23 as alternative to Na23 + p ⟶ He4 + Ne20 (Q = -4.839 MeV).
validation: missing Na23 + p ⟶ n + Mg23 as alternative to Na23 + p ⟶ Mg24 + 𝛾 (Q = -4.839 MeV).
[9]:
False
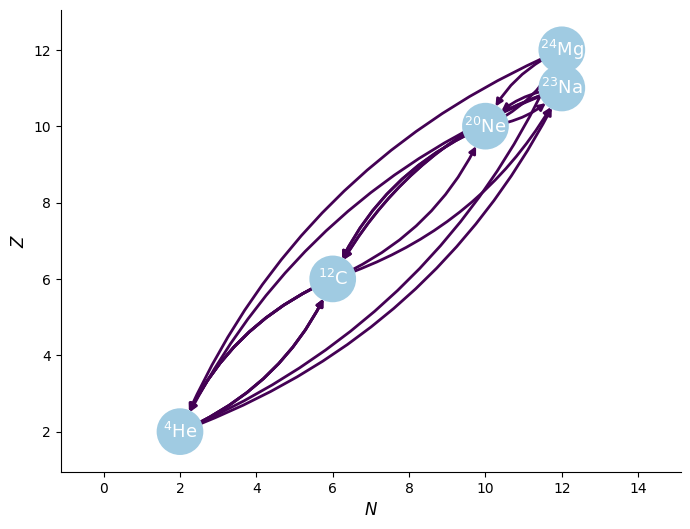
This now seems reasonable. The reactions that are missing are all endothermic.
[10]:
fig = rc.plot(curved_edges=True)
Finding duplicate links in a RateCollection
Sometimes we combine multiple sources of rates into a RateCollection or network, and we run the risk of having the same rate sequence duplicated, even though the rate itself may be different (i.e., a different source or tabulated vs. ReacLib).
Here we see how to check a RateCollection for duplicates.
We’ll start by recreating the electron-capture supernova network explored earlier.
First we get rates from ReacLib
[11]:
all_nuclei = ["p", "he4",
"ne20", "o20", "f20",
"mg24", "al27", "o16",
"si28", "s32", "p31"]
ecsn_rl_library = reaclib_library.linking_nuclei(all_nuclei,with_reverse=True)
Next we get some tabulated rates
[12]:
tl = pyna.TabularLibrary()
ecsn_tabular_library = tl.linking_nuclei(["f20", "o20", "ne20"])
Now we build a single Library using these two libraries
[13]:
complete_library = ecsn_rl_library + ecsn_tabular_library
If we print out the rates, we see that there are duplicates involving beta decays
[14]:
complete_library.get_rates()
[14]:
[O20 ⟶ F20 + e⁻ + 𝜈,
F20 ⟶ Ne20 + e⁻ + 𝜈,
Ne20 ⟶ He4 + O16,
Mg24 ⟶ He4 + Ne20,
Si28 ⟶ p + Al27,
Si28 ⟶ He4 + Mg24,
P31 ⟶ He4 + Al27,
S32 ⟶ p + P31,
S32 ⟶ He4 + Si28,
O16 + He4 ⟶ Ne20 + 𝛾,
Ne20 + He4 ⟶ Mg24 + 𝛾,
Mg24 + He4 ⟶ Si28 + 𝛾,
Al27 + p ⟶ Si28 + 𝛾,
Al27 + He4 ⟶ P31 + 𝛾,
Si28 + He4 ⟶ S32 + 𝛾,
P31 + p ⟶ S32 + 𝛾,
O16 + O16 ⟶ p + P31,
O16 + O16 ⟶ He4 + Si28,
Mg24 + He4 ⟶ p + Al27,
Al27 + p ⟶ He4 + Mg24,
Si28 + He4 ⟶ p + P31,
Si28 + He4 ⟶ O16 + O16,
P31 + p ⟶ He4 + Si28,
P31 + p ⟶ O16 + O16,
F20 ⟶ Ne20 + e⁻ + 𝜈,
F20 + e⁻ ⟶ O20 + 𝜈,
Ne20 + e⁻ ⟶ F20 + 𝜈,
O20 ⟶ F20 + e⁻ + 𝜈]
The find_duplicate_links() method will find these
[15]:
dupes = complete_library.find_duplicate_links()
dupes
[15]:
[[O20 ⟶ F20 + e⁻ + 𝜈, O20 ⟶ F20 + e⁻ + 𝜈],
[F20 ⟶ Ne20 + e⁻ + 𝜈, F20 ⟶ Ne20 + e⁻ + 𝜈]]
Here we see 2 pairs of duplicates. Looking at the first
[16]:
for rate in dupes[0]:
print(f"{rate.fname:20} {type(rate)}")
O20__F20__weak__wc12 <class 'pynucastro.rates.rate.ReacLibRate'>
O20__F20 <class 'pynucastro.rates.rate.TabularRate'>
We see that one is a ReacLibRate and the other is a TabularRate. In general, the TabularRate is preferred, since it has a density and temperature dependence and works at high densities.
We can use this information to remove the undesired rate from the RateCollection using the remove_rates() method.
[17]:
from pynucastro.rates import ReacLibRate
rates_to_remove = []
for d in dupes:
rates_to_remove += [r for r in d if isinstance(r, ReacLibRate)]
[18]:
for r in rates_to_remove:
complete_library.remove_rate(r)
Now we see only a single instance of these links
[19]:
complete_library.get_rates()
[19]:
[Ne20 ⟶ He4 + O16,
Mg24 ⟶ He4 + Ne20,
Si28 ⟶ p + Al27,
Si28 ⟶ He4 + Mg24,
P31 ⟶ He4 + Al27,
S32 ⟶ p + P31,
S32 ⟶ He4 + Si28,
O16 + He4 ⟶ Ne20 + 𝛾,
Ne20 + He4 ⟶ Mg24 + 𝛾,
Mg24 + He4 ⟶ Si28 + 𝛾,
Al27 + p ⟶ Si28 + 𝛾,
Al27 + He4 ⟶ P31 + 𝛾,
Si28 + He4 ⟶ S32 + 𝛾,
P31 + p ⟶ S32 + 𝛾,
O16 + O16 ⟶ p + P31,
O16 + O16 ⟶ He4 + Si28,
Mg24 + He4 ⟶ p + Al27,
Al27 + p ⟶ He4 + Mg24,
Si28 + He4 ⟶ p + P31,
Si28 + He4 ⟶ O16 + O16,
P31 + p ⟶ He4 + Si28,
P31 + p ⟶ O16 + O16,
F20 ⟶ Ne20 + e⁻ + 𝜈,
F20 + e⁻ ⟶ O20 + 𝜈,
Ne20 + e⁻ ⟶ F20 + 𝜈,
O20 ⟶ F20 + e⁻ + 𝜈]
and we can make a RateCollection now
[20]:
rc = pyna.RateCollection(libraries=[complete_library])